Introduction
Transcriptomics studies RNA in cells as a way of telling which parts of the genome are turned on and being actively expressed [1]. Comparing transcriptomes is useful in experiments to determine how gene expression changes between experimental conditions [4]. Some RNAs in the transcriptome will go on to become proteins and some will be active in their RNA form [2].
Microarrays measure the transcriptome by quantifying interactions of a pre-picked set of sequences. RNA sequencing is able to capture all RNAs produced in a cellular environment at a given moment for comparison of expression profiles [2].
In the case of the Human Protein Atlas RNA sequencing is able to indicate which tissues a gene is expressed in which can give insight on the role the gene plays in the body.
Microarrays measure the transcriptome by quantifying interactions of a pre-picked set of sequences. RNA sequencing is able to capture all RNAs produced in a cellular environment at a given moment for comparison of expression profiles [2].
In the case of the Human Protein Atlas RNA sequencing is able to indicate which tissues a gene is expressed in which can give insight on the role the gene plays in the body.
Results
Discussion
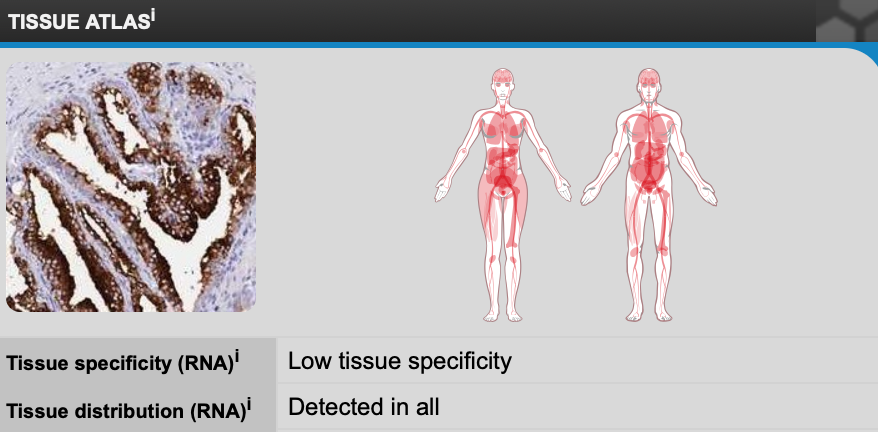
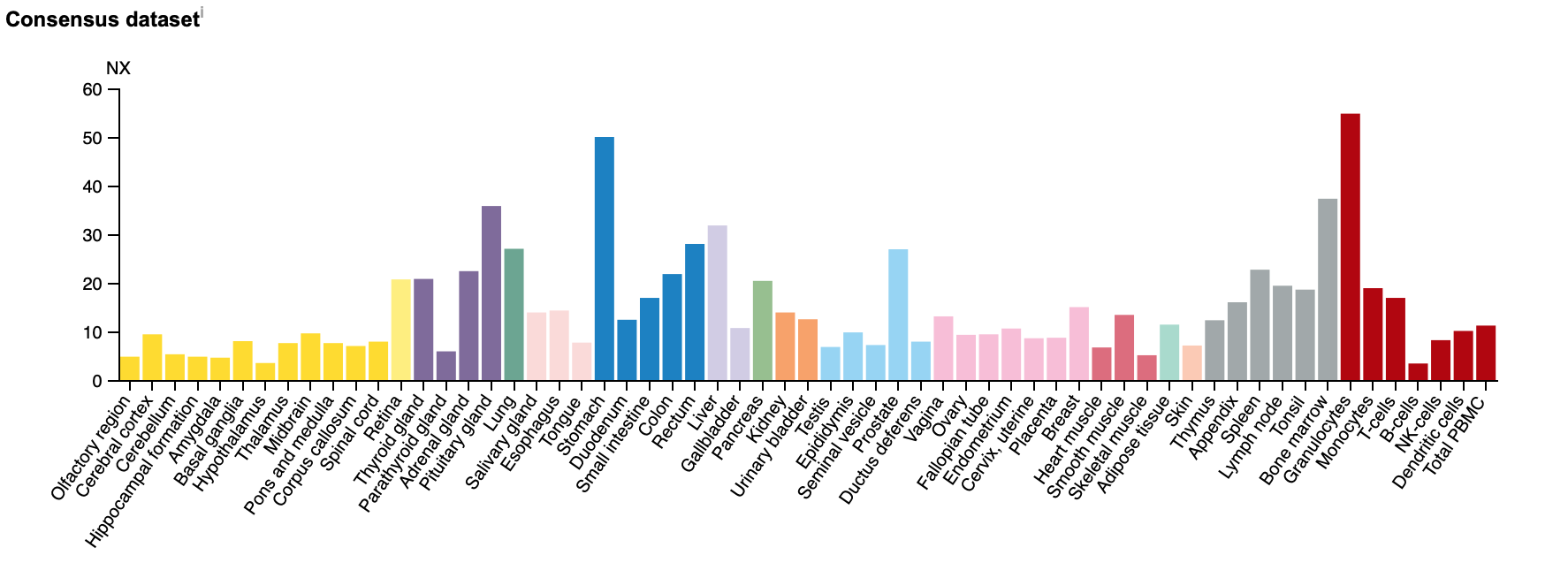
Some of the peaks in tissue specific RAB27A are very fitting for the symptoms of Griscelli Syndrome (GS). High expression in granulocytes, immune cells, show how in healthy individuals RAB27A function is important to cell function. Loss of function in those cells results in the immunodeficiency so well associated with GS Type II.
What is especially interesting is that RAB27A is expressed in most tissues in the body and yet mutations in the gene are associated with very specific symptoms in very specific tissues. One would think that loss of function in a gene expressed throughout the body would have far more spread and disastrous phenotypes. This lack of understanding of why the phenotypes of RAB27A mutants are so specific feeds into the gap in knowledge on GS.
What is especially interesting is that RAB27A is expressed in most tissues in the body and yet mutations in the gene are associated with very specific symptoms in very specific tissues. One would think that loss of function in a gene expressed throughout the body would have far more spread and disastrous phenotypes. This lack of understanding of why the phenotypes of RAB27A mutants are so specific feeds into the gap in knowledge on GS.
References
[1]Blackburn, Laura. “What Is Transcriptomics?” PHG Foundation, 18 Aug. 2017, www.phgfoundation.org/blog/what-is-transcriptomics.
[2]Lowe, Rohan, et al. “Transcriptomics Technologies.” PLOS Computational Biology, vol. 13, no. 5, 2017, doi:10.1371/journal.pcbi.1005457.
[3]“RAB27A.” RAB27A Protein Expression Summary - The Human Protein Atlas, www.proteinatlas.org/ENSG00000069974-RAB27A.
[4]“Transcriptomics.” Nature News, Nature Publishing Group, www.nature.com/subjects/transcriptomics.
[1]Blackburn, Laura. “What Is Transcriptomics?” PHG Foundation, 18 Aug. 2017, www.phgfoundation.org/blog/what-is-transcriptomics.
[2]Lowe, Rohan, et al. “Transcriptomics Technologies.” PLOS Computational Biology, vol. 13, no. 5, 2017, doi:10.1371/journal.pcbi.1005457.
[3]“RAB27A.” RAB27A Protein Expression Summary - The Human Protein Atlas, www.proteinatlas.org/ENSG00000069974-RAB27A.
[4]“Transcriptomics.” Nature News, Nature Publishing Group, www.nature.com/subjects/transcriptomics.
The web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison